Some Context
After reading this really good machine learning textbook by Brett Lantz, I was left wondering where I could use some of the techniques taught in the book for my personal use.
Personally, when I search for jobs on websites like Seek, I tend to narrow my search to a specific role. The dangers with this is that, in recent times, data roles can go by many names. Different companies may use a different title for essentially the same role. Searching can therefore become difficult because of this. Even if you do find a role that sounds right for you, you realise after reading the job description it isn’t!
Using a dataset available in Kaggle (Australian job listings data from Seek job board), I decided to deconstruct the job description, word by word, to build a Naive Bayes classifier that could predict which industry the job falls under.
Workflow
The workflow follows closely to what Brett Lantz uses in his textbook - so check the textbook out!
- First step - read in csv file into R. Note the encoding - the file had some uncommon characters that was making it difficult to use string functions.
data <- read_csv("seek_australia_sample.csv",locale = readr::locale(encoding = "windows-1252"))
## Parsed with column specification:
## cols(
## pageurl = col_character(),
## crawl_timestamp = col_character(),
## job_title = col_character(),
## category = col_character(),
## company_name = col_character(),
## city = col_character(),
## post_date = col_datetime(format = ""),
## job_description = col_character(),
## job_type = col_character(),
## job_board = col_character(),
## geo = col_character(),
## state = col_character(),
## salary_offered = col_character()
## )
data <- data %>%
select(job_description,category) %>%
filter(!is.na(job_description))- For the purposes of this blog, I decided to focus on the Healthcare & Medical industry. I also created a classifier for Banking and Financial Industry but the results were less than clearcut with jobs in the Information & Communication Technology and Insurance industry among others overlapping (as expected given the similarities between the industries). Healthcare & Medical appeared more insulated, although a few industries did encroach, in particular, Education and Community Services.
data_flagged <- data %>%
mutate(flag=ifelse(category %in% c("Healthcare & Medical"),"Yes","No")) %>%
mutate(no_chars=nchar(job_description))- The following utilises a text mining package named tm. More details of this package can be found in Text Mining Infrastructure in R, Feinerer I, Hornik K and Meyer D, Journal of Statistical Software, 2008, Vol. 25, pp. 1-54. I will not go into detail with the following code but you can consult the textbook or the Journal.
descriptions_corpus <- VCorpus(VectorSource(data_flagged$job_description)) ##supply job descriptions into VCorpus
descriptions_corpus_clean <- tm_map(descriptions_corpus,content_transformer(tolower)) ##text transformation - lowercase
descriptions_corpus_clean <- tm_map(descriptions_corpus_clean, removeNumbers) ##remove numbers
descriptions_corpus_clean <- tm_map(descriptions_corpus_clean,removeWords, stopwords()) ##list of stopwords listed in the tm package
descriptions_corpus_clean <- tm_map(descriptions_corpus_clean, removePunctuation)
descriptions_corpus_clean <- tm_map(descriptions_corpus_clean, stripWhitespace)
descriptions_dtm <- DocumentTermMatrix(descriptions_corpus_clean) ##this will create a sparse matrix with all the words found in the above having its own column with the number of its occurrence for each job description listed in the cell- We now create the training and test dataset, here I’ve used a 80:20 split (remember to set your seed to ensure reproducibility). Play around with the seed to ensure you have a proper representation of your industry in both datasets.
set.seed(1234501)
train_sample <- sample(4725,3780)
descriptions_dtm_train <- descriptions_dtm[train_sample,]
descriptions_dtm_test <- descriptions_dtm[-train_sample,]
descriptions_train_labels <- data_flagged[train_sample,]$flag
descriptions_test_labels <- data_flagged[-train_sample,]$flagprop.table(table(descriptions_train_labels))
## descriptions_train_labels
## No Yes
## 0.91719577 0.08280423
prop.table(table(descriptions_test_labels))
## descriptions_test_labels
## No Yes
## 0.92063492 0.07936508- Following is a wordcloud of those job descriptions marked as belonging to the Healthcare & Medical Industry with words that appear the most appearing bigger and bolder. Looks consistent with jobs in this industry?
yes <- subset(data_flagged,flag=="Yes")
wordcloud(yes$job_description,max.words = 100,scale = c(3, 0.5),random.order = FALSE)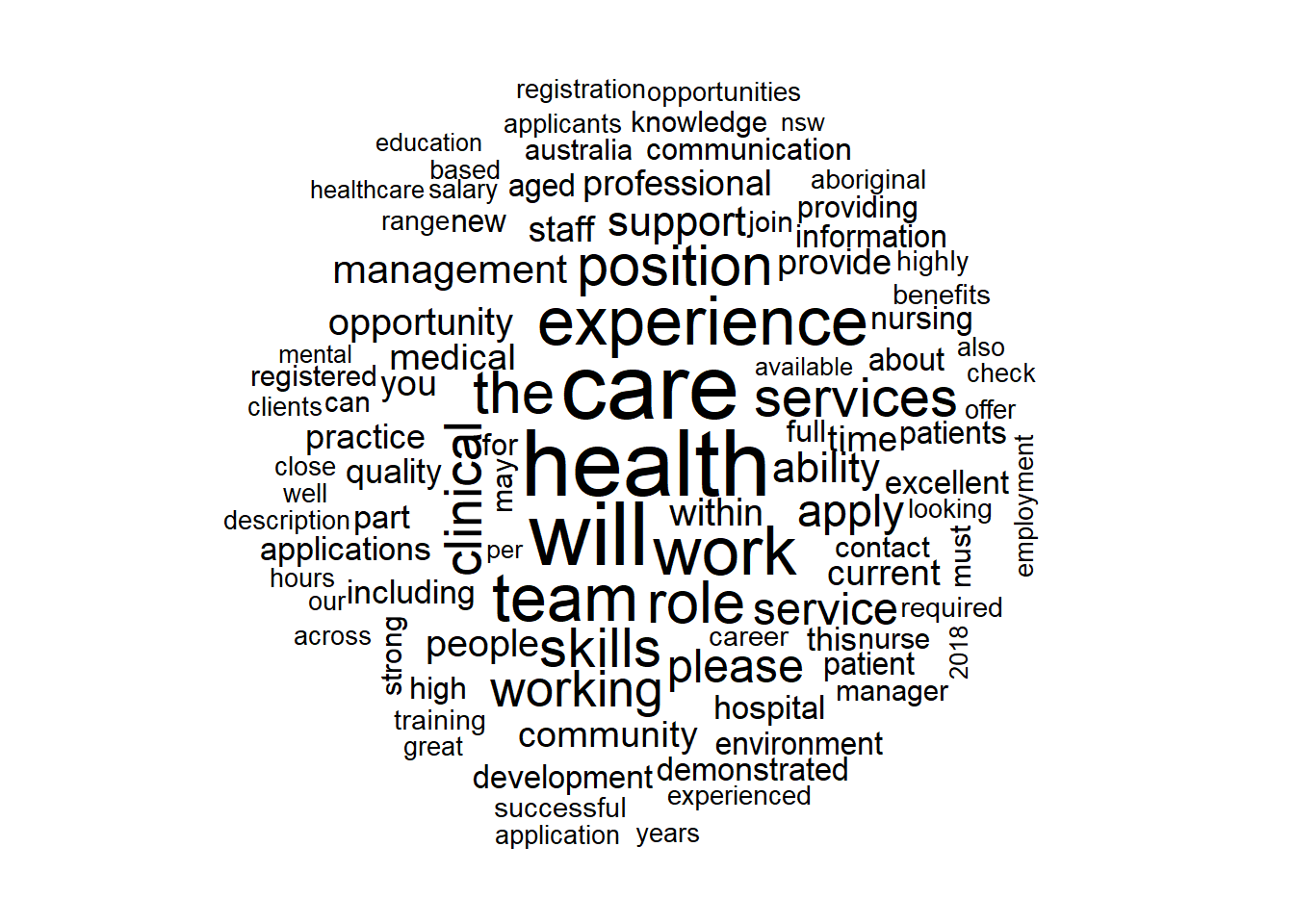
- We now need to reduce our test and training set by reducing the number of columns we have - at the moment it’s far too large! We do this by filtering them to only frequently occurring words (in my case, the word needs to appear in at least 60 job descriptions in the training dataset). Once this is done, our matrix needs to be transformed so that each cell represents an indicator (1 indicating that the word appears and a 0 meaning it doesn’t appear in the job description). This is because Naive Bayes classifier needs categorical features.
descriptions_freq_words <- findFreqTerms(descriptions_dtm_train, 60)
descriptions_dtm_freq_train <- descriptions_dtm_train[,descriptions_freq_words]
descriptions_dtm_freq_test <- descriptions_dtm_test[,descriptions_freq_words]
convert_counts <- function(x) {
x <- ifelse(x > 0, "Yes", "No")
}
descriptions_train <- as.data.frame(apply(descriptions_dtm_freq_train, MARGIN = 2,
convert_counts))
descriptions_test <- as.data.frame(apply(descriptions_dtm_freq_test, MARGIN = 2,
convert_counts))- We now train our dataset using the Naive Bayes classifier available in the e1071 package. Please ensure that the target labels (i.e. Yes/No) are set to a factor vector, otherwise the following code will generate an error.
descriptions_classifier <- naiveBayes(descriptions_train, as.factor(descriptions_train_labels),laplace=1)- Let’s now make predictions on the target dataset and check the performance of the classifier.
descriptions_classifier <- naiveBayes(descriptions_train, as.factor(descriptions_train_labels),laplace=1)
descriptions_test_pred <- predict(descriptions_classifier, descriptions_test, type = "raw")
descriptions_test_class <- predict(descriptions_classifier, descriptions_test)
descriptions_test_pred_yes <- as.data.frame(descriptions_test_pred) %>%
select(Yes)
CrossTable(descriptions_test_class, descriptions_test_labels,
prop.chisq = FALSE, prop.c = FALSE, prop.r = FALSE,
dnn = c('predicted', 'actual'))
##
##
## Cell Contents
## |-------------------------|
## | N |
## | N / Table Total |
## |-------------------------|
##
##
## Total Observations in Table: 945
##
##
## | actual
## predicted | No | Yes | Row Total |
## -------------|-----------|-----------|-----------|
## No | 813 | 17 | 830 |
## | 0.860 | 0.018 | |
## -------------|-----------|-----------|-----------|
## Yes | 57 | 58 | 115 |
## | 0.060 | 0.061 | |
## -------------|-----------|-----------|-----------|
## Column Total | 870 | 75 | 945 |
## -------------|-----------|-----------|-----------|
##
## The model was able to classify 92 percent of all the job descriptions as belonging to the Healthcare & Medical industry or not. Not bad! Or is it? Recall the distribution of our labels, we had Healthcare & Medical industry representing approximately 8% of our test dataset. If we had guessed that all labels in the response were not in the Healthcare & Medical industry, we would have also achieved 92 percent accuracy - without all this coding as well!
We want to remove this mere chance from our accuracy. Kappa refers to what’s known as Cohen’s Kappa statistic. The Kappa statistic adjusts the accuracy scores by accounting for the model being entirely correct by mere chance. Note that there are myriad of measures currently being used for model performance, with the Kappa statistic just being one of them.
library(irr)
## Loading required package: lpSolve
test_df <- as.data.frame(cbind(as.character(descriptions_test_class),descriptions_test_labels))
kappa2(test_df)
## Cohen's Kappa for 2 Raters (Weights: unweighted)
##
## Subjects = 945
## Raters = 2
## Kappa = 0.569
##
## z = 18
## p-value = 0The Kappa statistic of 56.9 percent suggests our model isn’t as good as we thought it initially was. However, the result isn’t too discouraging either - following off common interpretation of the Kappa statistic, the model is providing moderate agreement.
An important point when it comes to models and making predictions is that there’s a high element of subjectivity involved. For our purposes, we don’t need absolute perfection when determining what jobs we want to see (we can tolerate a bit of leeway), this would be another story if we were trying to detect cancer lumps!
- Now, imagine if we followed through with our model and had a system in place that picked up job descriptions through a job site API, we would be worried about the false negatives (17), jobs that were in the industry we were after but which we did not receive because our classifier said they weren’t! Rest assured, there are methodologies to skew the results so that false negatives are penalised and therefore reduce its occurrence - there’s one for decision trees in the C5.0 package.
Yikes, this blog is getting longer than I anticipated. I’ll wrap it up here. Following is a glimpse of the industries prominent in the false positives - notice the potential for overlaps between the top three industries. Full code available here.
as.data.frame(cbind(A=data_flagged[-train_sample,]$job_description,category=data_flagged[-train_sample,]$category,descriptions_test_class,descriptions_test_pred_yes)) %>%
filter(descriptions_test_class=="Yes") %>%
filter(!(category %in% c("Healthcare & Medical"))) %>%
group_by(category) %>%
tally() %>%
arrange(desc(n)) %>%
filter(n>1)
## # A tibble: 5 x 2
## category n
## <fct> <int>
## 1 Community Services & Development 18
## 2 Education & Training 18
## 3 Government & Defence 6
## 4 Administration & Office Support 4
## 5 Information & Communication Technology 2